Cython
Please open notebook rsepython-s6r4.ipynb
Cython can be viewed as an extension of Python where variables and functions are annotated with extra information, in particular types. The resulting Cython source code will be compiled into optimised C or C++ code, and thereby yielding substantial speed-up of slow Python code. In other word, Cython provides a way of writing Python with comparable performance to that of C/C++.
Start Coding in Cython
Cython code must, unlike Python, be compiled. This happens in the following stages:
- The cython code in
.pyxfile will be translated to aCfile. - The
Cfile will be compiled by a C compiler into a shared library, which will be directly loaded into Python.
In the Jupyter notebook, everything is a lot easier. One need only to load Cython extension (%load_ext Cython) at the beginning and put %%cython mark in front of cells of cython code. Cells with cython mark will be treated as a .pyx code and consequently, compiled into C.
For details, please see Building Cython Code.
Pure python Mandelbrot set:
xxmin = -1.5
ymin = -1.0
xmax = 0.5
ymax = 1.0
resolution = 300
xstep = (xmax - xmin) / resolution
ystep = (ymax - ymin) / resolution
xs = [(xmin + (xmax - xmin) * i / resolution) for i in range(resolution)]
ys = [(ymin + (ymax - ymin) * i / resolution) for i in range(resolution)]
def mandel(position, limit=50):
value = position
while abs(value) < 2:
limit -= 1
value = value**2 + position
if limit < 0:
return 0
return limit
Compiled by Cython:
%load_ext Cython
%%cython
def mandel_cython(position, limit=50):
value = position
while abs(value) < 2:
limit -= 1
value = value**2 + position
if limit < 0:
return 0
return limit
Let’s verify the result
from matplotlib import pyplot as plt
%matplotlib inline
f, axarr = plt.subplots(1, 2)
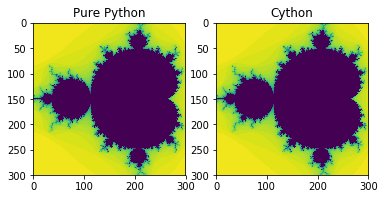
axarr[0].imshow([[mandel(complex(x, y)) for x in xs] for y in ys], interpolation='none')
axarr[0].set_title('Pure Python')
axarr[1].imshow([[mandel_cython(complex(x, y)) for x in xs] for y in ys], interpolation='none')
axarr[1].set_title('Cython')
Text(0.5,1,’Cython’)
%timeit [[mandel(complex(x, y)) for x in xs] for y in ys] # pure python
%timeit [[mandel_cython(complex(x, y)) for x in xs] for y in ys] # cython
1 loop, best of 3: 711 ms per loop
1 loop, best of 3: 474 ms per loop
We have improved the performance of a factor of 1.5 by just using the cython compiler, without changing the code!
Cython with C Types
But we can do better by telling Cython what C data type we would use in the code. Note we’re not actually writing C, we’re writing Python with C types.
typed variable
%%cython
def var_typed_mandel_cython(position, limit=50):
cdef double complex value # typed variable
value = position
while abs(value) < 2:
limit -= 1
value = value**2 + position
if limit < 0:
return 0
return limit
typed function + typed variable
%%cython
cpdef call_typed_mandel_cython(double complex position,
int limit=50): # typed function
cdef double complex value # typed variable
value = position
while abs(value)<2:
limit -= 1
value = value**2 + position
if limit < 0:
return 0
return limit
performance of one number:
# pure python
%timeit a = mandel(complex(0, 0))
100000 loops, best of 3: 14.5 µs per loop
# primitive cython
%timeit a = mandel_cython(complex(0, 0))
100000 loops, best of 3: 9 µs per loop
# cython with C type variable
%timeit a = var_typed_mandel_cython(complex(0, 0))
100000 loops, best of 3: 6.45 µs per loop
# cython with typed variable + function
%timeit a = call_typed_mandel_cython(complex(0, 0))
100000 loops, best of 3: 3.23 µs per loop
Cython with numpy ndarray
You can use NumPy from Cython exactly the same as in regular Python, but by doing so you are losing potentially high speedups because Cython has support for fast access to NumPy arrays.
import numpy as np
ymatrix, xmatrix = np.mgrid[ymin:ymax:ystep, xmin:xmax:xstep]
values = xmatrix + 1j * ymatrix
%%cython
import numpy as np
cimport numpy as np
cpdef numpy_cython_1(np.ndarray[double complex, ndim=2] position,
int limit=50):
cdef np.ndarray[long,ndim=2] diverged_at
cdef double complex value
cdef int xlim
cdef int ylim
cdef double complex pos
cdef int steps
cdef int x, y
xlim = position.shape[1]
ylim = position.shape[0]
diverged_at = np.zeros([ylim, xlim], dtype=int)
for x in xrange(xlim):
for y in xrange(ylim):
steps = limit
value = position[y,x]
pos = position[y,x]
while abs(value) < 2 and steps >= 0:
steps -= 1
value = value**2 + pos
diverged_at[y,x] = steps
return diverged_at
Note the double import of numpy: the standard numpy module and a Cython-enabled version of numpy that ensures fast indexing of and other operations on arrays. Both import statements are necessary in code that uses numpy arrays. The new thing in the code above is declaration of arrays by np.ndarray.
%timeit data_cy = [[mandel(complex(x, y)) for x in xs] for y in ys] # pure python
1 loop, best of 3: 692 ms per loop
%timeit data_cy = [[call_typed_mandel_cython(complex(x, y)) for x in xs] for y in ys] # typed cython
1 loop, best of 3: 201 ms per loop
%timeit numpy_cython_1(values) # ndarray
10 loops, best of 3: 192 ms per loop
A trick of using np.vectorize
numpy_cython_2 = np.vectorize(call_typed_mandel_cython)
%timeit numpy_cython_2(values) # vectorize
1 loop, best of 3: 188 ms per loop
Calling C functions from Cython
Example: compare sin() from Python and C library
%%cython
import math
cpdef py_sin():
cdef int x
cdef double y
for x in range(1e7):
y = math.sin(x)
%%cython
from libc.math cimport sin as csin # import from C library
cpdef c_sin():
cdef int x
cdef double y
for x in range(1e7):
y = csin(x)
%timeit [math.sin(i) for i in range(int(1e7))] # python
1 loop, best of 3: 2.21 s per loop
%timeit py_sin() # cython call python library
1 loop, best of 3: 1.62 s per loop <?span>
%timeit c_sin() # cython call C library
100 loops, best of 3: 6.24 ms per loop